In summary
- The researchers presented Delphi-2M in nature, an AI that predicts the risk of more than 1,000 diseases up to 20 years.
- The model beat the unique disease tools, predicting comorbidities and generating synthetic health trajectories from medical records.
- Trained in the Biobank of the United Kingdom and validated in 1.9 million Danish health records, Delphi-2M is promising but faces biases, privacy and display obstacles.
Researchers have created an AI system that predicts their risk of developing more than 1,000 diseases up to 20 years before the symptoms appear, according to a study published in Nature this week.
The model, called Delphi-2M, reached 76% accuracy for short-term health predictions and maintained a 70% accuracy even when a decade was forecast in the future.
He surpassed existing unique disease risk calculators while simultaneously evaluating the risks throughout the spectrum of human diseases.
“The progression of human disease through age is characterized by health periods, episodes of acute disease and also chronic weakening, often manifesting as comorbidity groups,” the researchers wrote. “Few algorithms are able to predict the complete spectrum of human disease, which recognizes more than 1,000 diagnoses at the upper level of the international classification of diseases, the tenth review coding system (ICD-10).”
The system learned these patterns of 402,799 Biobank participants in the United Kingdom, then demonstrated its temper in 1.9 million Danish health records without any additional training.
Before starting to rub your hands with the idea of your own medical predictor, can you try Delphi-2M yourself? Not quite.
The trained model and its weights are blocked behind the controlled access procedures of the United Kingdom biobank, which means only researchers. The code base to train its own version is in Github under a MIT license, so you could technically build your own model, but you would need access to mass medical data sets to work.
For now, this remains a research tool, not a consumption application.
Behind the curtain
Technology works by treating medical records as sequences, as well as the text of the chatgpt processes.
Each diagnosis, registered with the age that occurred for the first time, becomes a file. The model reads this medical “language” and predicts what is coming later.
With adequate information and training, you can predict the following token (in this case, the next disease) and the estimated time before that “token” is generated (how long until the most likely set of events occurs).
For a 60-year-old with diabetes and high blood pressure, Delphi-2M could forecast a 19-time risk of pancreas cancer. Add a diagnosis of pancreatic cancer to that history, and the model calculates the risk of mortality that jumps almost ten thousand times.
The transformer architecture behind Delphi-2M represents each person’s health trip as a timeline of diagnostic codes, lifestyle factors such as smoking and BMI, and demographic data. Padded tokens “without event” fill the gaps among medical visits, teaching the model that the simple passage of time changes the reference risk.
This is also similar to how normal LLMs can understand the text, even if they lose some words or even sentences.
When tested against established clinical tools, Delphi-2M coincided or exceeded its performance. For the prediction of cardiovascular disease, it achieved an AUC of 0.70 compared to 0.69 for self -process and 0.71 for Qrisk3. For dementia, it reached 0.81 versus 0.81 for UKBDRS. The key difference: these tools predict unique conditions. Delphi-2M evaluates everything at once.
Beyond individual predictions, the system generates whole synthetic health trajectories.
From the 60 -year data, it can simulate thousands of possible health futures, producing disease load estimates at the precise population level for statistical margins. A synthetic data set trained a secondary Delphi model that reached an accuracy of 74%, only three percentage points below the original.
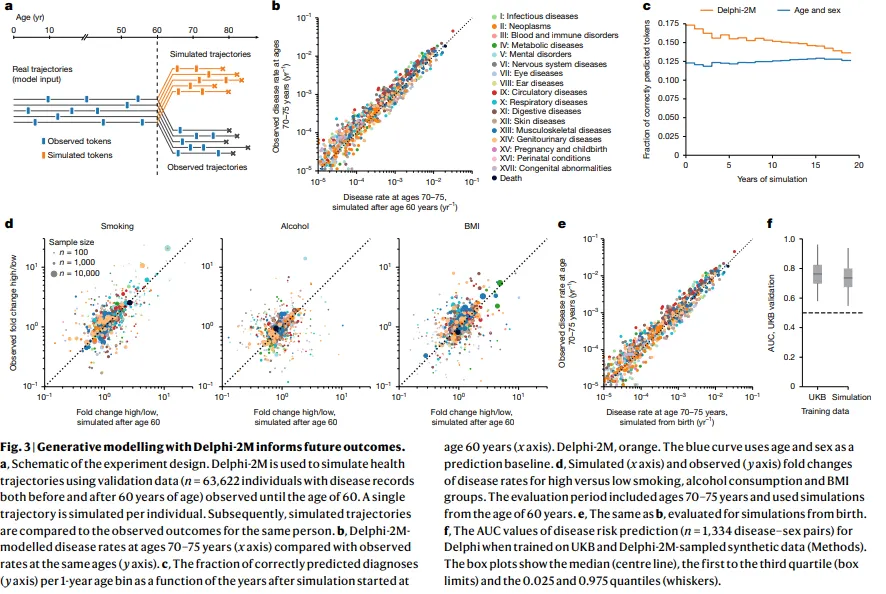
The model revealed how diseases influence each other over time. Cancers increased the risk of mortality with a “half -life” of several years, while the effect of septicemia fell sharply, returning to the nearby line in a matter of months. Mental health conditions showed persistent group effects, with a diagnosis that strongly predicts others in that category years later.
Limitations
The system has limits. Its 20-year predictions fall to around 60-70% precision in general, but things will depend on the type of disease and conditions that try to analyze and forecast.
“For 97% of the diagnoses, the AUC was greater than 0.5, which indicates that the vast majority followed the patterns with at least one partial predictability,” says the study, and added later that “the average AUC values of Delphi-2M decreases from an average of 0.76 to 0.70 after 10 years”, and that “it is the first year of samples, they are on average of 17% of the disease. late.”
In other words, this model is quite good to predict things in relevant scenarios, but much can change in 20 years, so it is not Nostradamus.
Rare diseases and highly environmental conditions are more difficult to forecast. The demographic bias of the United Kingdom biobank, mostly white, educated, relatively healthy volunteers, includes a bias that researchers recognize that the needs address.
Danish validation revealed another limitation: Delphi-2M learned some specific peculiarities of data collection of the United Kingdom. Diseases registered mainly in hospital environments appeared artificially inflated, contradicting the data recorded by the Danish people.
The model predicted septicemia eight times the normal rate for any person with previous hospital data, partly because 93% of the biobanco septicemia diagnoses of the United Kingdom come from hospital records.
The researchers trained Delphi-2M using a modified GPT-2 architecture with 2.2 million parameters, Tiny compared to modern but sufficient language models for medical prediction. The key modifications included the continuous coding of age instead of discrete position markers and an exponential waiting time model to predict when the events would occur, not only what would happen.
Each health career in training data contained an average of 18 disease tokens that cover birth until 80 years. Sex, BMI categories, state of smoking and alcohol consumption added context.
The model learned to weigh these factors automatically, discovering that obesity increased the risk of diabetes while smoking elevated cancer probabilities, relationships that medicine has established for a long time but that arose without explicit programming. It is really a LLM for health conditions.
For the clinical deployment, several obstacles remain.
The model needs validation in more diverse populations, for example, lifestyles and habits of people in Nigeria, China and the United States can be very different, which makes the model less precise.
In addition, privacy concerns about the use of detailed health records require careful management. Integration with existing health systems raises technical and regulatory challenges.
But potential applications cover from identifying detection candidates that do not comply with age -based criteria until they model population health interventions. Insurance companies, pharmaceutical companies and public health agencies can have obvious interests.
Delphi-2M joins a growing family of medical models based on transformers. Some examples Include Harvard’s PDGRAPHER tool to predict gene drug combinations that could reverse diseases such as Parkinson’s or Alzheimer’s, a LLM specifically trained in protein connections, the Google alfagenoma model trained in DNA pairs and others.
What makes Delphi-2M so interesting and different is its wide range of action, the great amplitude of covered diseases, their long prediction horizon and their ability to generate realistic synthetic data that preserve statistical relationships while protecting individual privacy.
In other words: “How long do I have?” Soon it can be less a rhetorical question and more a predictable data point.
Usually intelligent Information sheet
A weekly journey of gene, a generative model.


